MAJOR UPDATE
As of Fall 2021, this wiki has been discontinued and is no longer being actively developed.
All updated materials and announcements for the QCBS R Workshop Series are now housed on the QCBS R Workshop website. Please update your bookmarks accordingly to avoid outdated material and/or broken links.
Thank you for your understanding,
Your QCBS R Workshop Coordinators.
QCBS R Workshops
This series of 10 workshops walks participants through the steps required to use R for a wide array of statistical analyses relevant to research in biology and ecology. These open-access workshops were created by members of the QCBS both for members of the QCBS and the larger community.
The content of this workshop has been peer-reviewed by several QCBS members. If you would like to suggest modifications, please contact the current series coordinators, listed on the main wiki page
IMPORTANT NOTICE: MAJOR UPDATES
March 2021 update: This wiki has been discontinued and is no longer being actively developed or updated. Updated materials for the QCBS R Workshop Series are now housed on the QCBS R Workshop GitHub page.
Available materials include;
- The Rmarkdown presentation for this workshop;
- An R script that follows the presentation;
- Written materials that accompany the presentation in bookdown format.
Workshop 10: Advanced multivariate analyses
Developed by: Monica Granados, Emmanuelle Chrétien, Bérenger Bourgeois, Amanda Winegardner and Xavier Giroux-Bougard. (Material in R script adapted from: Borcard, Gillet & Legendre (2011). Numerical Ecology with R. Springer New York.)
Summary: In this workshop, you will learn how to perform advanced multivariate analyses on community data. This workshop concentrates on constrained methods such as redundancy analysis (RDA), multivariate regression tree (MRT) and linear discriminant analysis (LDA) to explore how environmental variables may be driving patterns in species assemblage across sites.
Link to new Rmarkdown presentation
Link to old Prezi presentation
Download the R script, packages and data required for this workshop:
Make sure to load the following packages (see how in the R script):
- mvpart
- MVPARTwrap
- rdatest
- | Load the required packages and functions
install.packages("vegan") install.packages("mvpart") install.packages("labdsv") install.packages("plyr") install.packages("MASS") # For the two following packages, upload the file provided on the wiki page. # To do so, go to Packages tab on the bottom right panel of R Studio # Click on Install Packages # Choose to install from Package Archive file and upload these two files install.packages("MVPARTwrap") install.packages("rdaTest") library(vegan) library(mvpart) library(MVPARTwrap) library(rdaTest) library(labdsv) library(plyr) library(MASS)
Introduction to advanced multivariate analyses
The previous workshop presented the basics of multivariate analyses:
- how to choose appropriate distance metrics and transformations
- hierarchical clustering
- unconstrained ordinations
- Principal component analysis
- Principal coordinate Analysis
- Correspondence analysis
- Nonmetric multidimensional scaling
The present workshop builds on this knowledge, and will focus on constrained analyses. All the methods overviewed during the introductory workshop allowed to find patterns in the community composition data or in the descriptors, but not to explore how environmental variables could be driving these patterns. With constrained analyses, such as redundancy analysis (RDA), linear discriminant analysis (LDA) and multivariate regression tree (MRT), one can describe and predict relationships between community composition data and environmental variables.
Getting started with data
We will continue to use the Doubs river datasets for this workshop. “DoubsSpe.csv” is a data frame of fish community data where the first column contains site names from 1 to 30 and the remaining columns are fish taxa. The taxa columns are populated by fish abundance data (counts). “DoubsEnv.csv” is a data frame of environmental data for the same sites contained in the fish community data frame. Again, the first column contains site names from 1 to 30. The remaining columns contain measurements for 11 abiotic variables. Note that data used in ordination analyses is generally in wide-format.
- | Load the Doubs Species and Environmental Data
#Species community data frame (fish abundance): “DoubsSpe.csv” spe<- read.csv(file.choose(), row.names=1) spe<- spe[-8,] #Site number 8 contains no species and so row 8 (site 8) is removed. Be careful to #only run this command line once as you are overwriting "spe" each time. #Environmental data frame: “DoubsEnv.csv” env<- read.csv(file.choose(), row.names=1) env<- env[-8,] #Remove corresponding abiotic data for site 8 (since removed from fish data). #Again, be careful to only run the last line once.
1. Explore and prepare the data
1.1 Species data
We can use summary functions to explore the “spe” data (fish community data) and discover things like the dimensions of the matrix, column headings and summary statistics for the columns. This is a review from Workshop 2.
- | Explore DoubsSpe
names(spe) #see names of columns in spe dim(spe) #dimensions of spe; number of columns and rows str(spe) #displays internal structure of objects head(spe) #first few rows of the data frame summary(spe) #summary statistics for each column; min value, median value, max value, mean value etc.
Look at the species’ distribution frequencies.
- | Species distribution of DoubsSpe data
#Species distribution (ab <- table(unlist(spe))) #note that when you put an entire line of code in brackets like this, the output for that operation is displayed right away in the R console barplot(ab, las=1, xlab="Abundance class", ylab="Frequency", col=grey(5:0/5))
 Can see that there is a high frequency of zeros in the abundance data.
Can see that there is a high frequency of zeros in the abundance data.
See how many absences there are in the fish community data.
- | Absences in fish data
sum(spe==0)
Look at the proportion of zeros in the fish community data.
- | Proportion of zeros
sum(spe==0)/(nrow(spe)*ncol(spe))
The proportion of zeros in the dataset is ~0.5.
This is high but not uncommon for species abundance data. However, in order to avoid the use of double-zeros as indications of resemblance among sites, we will apply a transformation to the species data. Legendre and Gallagher (2001) proposed five pre-transformations of the species data, four of them being available in vegan in the function decostand().
The Hellinger transformation will be applied to the fish data. It expresses abundances as the square-root of their relative abundance at each site (Borcard et al. 2011).
- | Hellinger transformation
spe.hel <- decostand(spe, method="hellinger") # you can also use method="hell"
1.2 Environmental data
Explore the environmental data and create a panel of plots to compare collinearity:
- | Explore Doubs Env data
names(env) dim(env) str(env) head(env) summary(env) pairs(env, main="Bivariate Plots of the Environmental Data" )
In this case, the environmental data (explanatory variables) are all in different units and need to be standardized prior to computing distance measures to perform most ordination analyses. Standardize the environmental data (11 variables) using the function decostand() in vegan.
- | Decostand
env.z <- decostand(env, method="standardize") apply(env.z, 2, mean) # the data are now centered (means~0) apply(env.z, 2, sd) # the data are now scaled (standard deviations=1)
2. Canonical analyses
Described first by Rao (1964), canonical analysis is a generic term that for several types of statistical analyses sharing a common goal; to identify the relationship between a multivariate response table (matrix Y, generally describing the species composition of communities) and a multivariate explanatory table (matrix X, generally containing environmental descriptors) by combining ordination and regression concepts. Canonical analyses allow users to test ecological hypothesis concerning the environmental drivers of species composition. Among the diversity of canonical analysis, we will mainly focus here on Redundancy Analysis (RDA).
2.1 Redundancy analysis (RDA)
Redundancy Analysis is a direct extension of multiple regression, as it models the effect of an explanatory matrix X (n x p) on a response matrix Y (n x m). This is done by preforming an ordination of Y to obtain ordination axes that are linear combinations of the variables in X. In RDA, ordination axes are calculating from a PCA of a matrix Yfit, computed by fitting the Y variables to X by multivariate linear regression. Note that the explanatory variables in X can be quantitative, qualitative or binary variables. Prior to RDA, explanatory variables in Y must be centered, standardized (if explanatory variables are not dimensionally homogeneous, i.e. in different units), transformed (to limit the skew of explanatory variables) or normalized (to linearize relationships) following the same principles as in PCA. Collinearity between the X variables should also be reduced before RDA.
In order to obtain the best model of RDA, explanatory variables can be selected by forward, backward or stepwise selection that remove non-significant explanatory variables.
RDA involves two computational steps. In the first step, a matrix of fitted values Yfit is calculated trough the linear equation: Yfit = X[X'X]-1 [X'Y]
In the second step, a PCA of the fitted matrix Yfit is calculated (see equations in the figure below) and produce the canonical eigenvalues and eigenvectors together with the matrix Z containing the canonical axes. These canonical axes correspond to linear combinations of the explanatory variables in X. This linearity of the combinations of the X variables is a fundamental property of RDA. In the analysis of community composition, these canonical axes are interpreted as complex environmental gradients.
Redundancy analysis as a two-step process (from Legendre and Legendre 2012) Several statistics can be computed from RDA:
- The R² measures the strength of the canonical relationship between Y and X by calculating the proportion of the variation of Y explained by the variables in X, - The adjusted R² also measures the strength of the relationship between Y and X, but applies a correction of the R² to take into account the number of explanatory variables, - The F-statistic corresponds to an overall test of significance of an RDA by comparing the computed model to a null model. This test is based on the null hypothesis that the strength of the linear relationship calculated by the R² is not larger than the value that would be obtained for unrelated Y and X matrices of the same size. Note that F-statistics can also be used to sequentially test the significance of each canonical axis.
In R, RDA can be computed using the function rda from package vegan, as follows:
- | rda() in vegan for RDA
#Preparing the data prior to RDA env.z <- subset(env.z, select = -das) # remove the "distance from the source" variable #Running the RDA ?rda spe.rda <- rda(spe.hel~., data=env.z) ### Extract the results summary(spe.rda, display=NULL) #The results are called using summary: summary(spe.rda, display=NULL) #display = NULL optional
The summary of the output looks like this:
These results contain the proportion of variance of Y explained by the X variables (constrained proportion, 73.41% here), the unexplained variance of Y (unconstrained proportion, 26.59% here) and then summarize the eigenvalues, the proportions explained and the cumulative proportion of each canonical axis (each canonical axis = each constraining variable, in this case, the environmental variables from env).
To select the significant explanatory variables, you can then perform a forward selection (or backwards or stepwise), using the ordiR2step() function (or using the forward.sel function of package packfor):
- | ordiR2step() for forward selection
### Select the significant explanatory variables by forward selection ?ordiR2step ordiR2step(rda(spe.hel~1, data=env.z), scope= formula(spe.rda), direction= "forward", R2scope=TRUE, pstep=1000) env.signif <- subset(env.z, select = c("alt", "oxy", "dbo"))
The resulting output reads: rda(formula = spe.hel ~ alt + oxy + dbo, data = env.z) with the proportion of variation explained by the three constraining variables being 0.59. So in this case, only three variables are retained by the forward selection, i.e. alt, oxy and dbo. These three variables can be placed in a new data frame env.signif to perform the new RDA including only significant X variables:
- | RDA with the two retained variables
spe.rda.signif <- rda(spe.hel~., data=env.signif) summary(spe.rda.signif, display=NULL)
The explanatory variables (altitude, oxygen and biological oxygen demand) now explain 59% of the variance in Y (species). The adjusted R² of this RDA is calculated using the function RsquareAdj:
- | adjusted R2
(R2adj <- RsquareAdj(spe.rda.signif)$adj.r.squared) #Here the strength of the relationship between X and Y corrected for the number of X variables is 0.54.
The significance of the model and of each canonical axis can be tested using the function anova (note this is different from retaining significant variables as was done with forward selection, now we're testing the significance of the RDA axes):
- | anova.cca for testing the significance of axes
?anova.cca anova.cca(spe.rda.signif, step=1000) anova.cca(spe.rda.signif, step=1000, by="axis") #In this case, the RDA model is highly significant (p=0.001) as well as all three canonical axes.
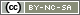
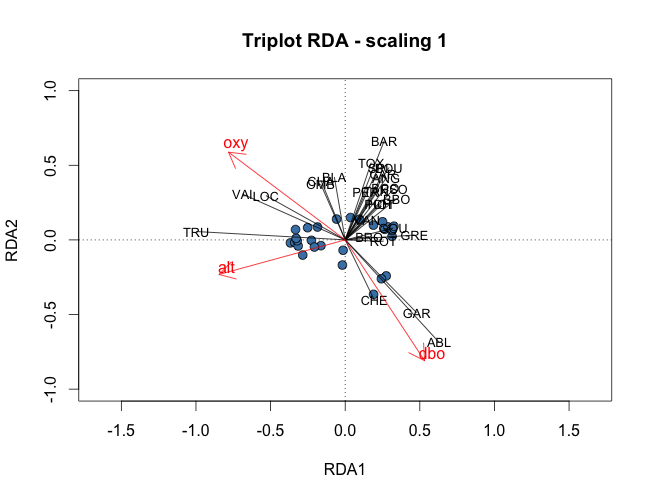
To visualize the results of the RDA, triplots can be drawn using the plot(). Note that as in PCA, users can create scaling 1 and scaling 2 triplots. In scaling 1, distance among objects approximate their Euclidean distances while in scaling 2, angles between variables X and Y reflect their correlation. Thus, scaling 1 triplots can be used to interpret distances among objects and scaling 2 triplots to interpret the relationships between X and Y. To plot the scaling 1 triplots of the RDA, the following code can be used:
- | plot RDAs
#Quick plots scaling 1 and 2 windows() plot(spe.rda.signif, scaling=1, main="Triplot RDA (scaling 1)") windows() plot(spe.rda.signif, scaling=2, main="Triplot RDA (scaling 2)") #Advanced plots scaling 1 windows() plot(spe.rda.signif, scaling=1, main="Triplot RDA - scaling 1", type="none", xlab=c("RDA1"), ylab=c("RDA2"), xlim=c(-1,1), ylim=c(-1,1)) points(scores(spe.rda.signif, display="sites", choices=c(1,2), scaling=1), pch=21, col="black", bg="steelblue", cex=1.2) arrows(0,0, scores(spe.rda.signif, display="species", choices=c(1), scaling=1), scores(spe.rda.signif, display="species", choices=c(2), scaling=1), col="black",length=0) text(scores(spe.rda.signif, display="species", choices=c(1), scaling=1), scores(spe.rda.signif, display="species", choices=c(2), scaling=1), labels=rownames(scores(spe.rda.signif, display="species", scaling=1)), col="black", cex=0.8) arrows(0,0, scores(spe.rda.signif, display="bp", choices=c(1), scaling=1), scores(spe.rda.signif, display="bp", choices=c(2), scaling=1), col="red") text(scores(spe.rda.signif, display="bp", choices=c(1), scaling=1)+0.05, scores(spe.rda.signif, display="bp", choices=c(2), scaling=1)+0.05, labels=rownames(scores(spe.rda.signif, display="bp", choices=c(2), scaling=1)), col="red", cex=1) #Advanced plots scaling 2 windows() plot(spe.rda.signif, scaling=2, main="Triplot RDA - scaling 2", type="none", xlab=c("RDA1"), ylab=c("RDA2"), xlim=c(-1,1), ylim=c(-1,1)) points(scores(spe.rda.signif, display="sites", choices=c(1,2), scaling=2), pch=21, col="black", bg="steelblue", cex=1.2) arrows(0,0, scores(spe.rda.signif, display="species", choices=c(1), scaling=2)*2, scores(spe.rda.signif, display="species", choices=c(2), scaling=2)*2, col="black",length=0) text(scores(spe.rda.signif, display="species", choices=c(1), scaling=2)*2.1, scores(spe.rda.signif, display="species", choices=c(2), scaling=2)*2.1, labels=rownames(scores(spe.rda.signif, display="species", scaling=2)), col="black", cex=0.8) arrows(0,0, scores(spe.rda.signif, display="bp", choices=c(1), scaling=2), scores(spe.rda.signif, display="bp", choices=c(2), scaling=2), col="red") text(scores(spe.rda.signif, display="bp", choices=c(1), scaling=2)+0.05, scores(spe.rda.signif, display="bp", choices=c(2), scaling=2)+0.05, labels=rownames(scores(spe.rda.signif, display="bp", choices=c(2), scaling=2)), col="red", cex=1)
And the final triplots would look like this:
Challenge 1: Run an RDA of the mite environmental variables constraining the mite species abundances. Use:
- | Load the mite data
#Load the mite species and environmental data from vegan package data(mite) mite.spe<-mite mite.spe.hel <- decostand(mite.spe, method="hellinger") data(mite.env)
What are the significant explanatory variables? How much of the variation in species data is explain by significant explanatory variables (i.e. those that were selected)? What are the significant axes? What group(s) of sites can you identify? What species are related to each group(s) of sites?
Challenge 1: Solution
2.2 Partial RDA
Partial RDA is a special case of RDA in which the response variables Y are related to explanatory variables X in the presence of additional explanatory variables, W, called covariables. As in partial linear regression, the linear effect of X variables on the Y variables are adjusted for the effects of the covariables W. For this, a RDA of the covariables W on the response variables Y is first performed. The residuals of this RDA are then extracted, i.e. a matrix Yres|W containing the Y response variables in which the effect of W were removed. The partial RDA finally correspond to the RDA of X on Yres|W. All statistics previously presented for RDA also apply for partial RDA.
Partial RDA is thus a powerful tool when users what to assess the effect of environmental variables on species composition while taking into account the species variation due to other environmental variables with no interest. It can also be used to control for well-known linear effects, isolate the effect of a single explanatory variable or analyse related samples. In the example below, we will assess the effect of water chemistry on fish species abundances partialling out the effect of physiography.
In R, partial RDA is performed in the same way as RDA using rda() with the addition of a condition term:
- | Partial RDA with rda()
#Divide the env2 dataframe into two dataframes: envtopo <- env[, c(1:3)] # Physiography : explanatory dataset 1 names(envtopo) envchem <- env[, c(4:10)] # Water quality : explanatory dataset 2 names(envchem) #Run the partial RDA spechem.physio=rda(spe.hel, envchem, envtopo) summary(spechem.physio, display=NULL) #or spechem.physio2=rda(spe.hel ~ pH + dur + pho + nit + amm + oxy + dbo + Condition(alt + pen + deb), data=env) #Extract the results summary(spechem.physio, display=NULL) #Calculate the adjusted R2 of the partial RDA (R2adj <- RsquareAdj(spechem.physio)$adj.r.squared) #Test the significance of the axes in partial RDA anova.cca(spechem.physio, step=1000) anova.cca(spechem.physio2, step=1000, by="axis") #Construct the triplots #Scaling 1 windows(title="Partial RDA scaling 1") plot(spechem.physio, scaling=1, main="Triplot partial RDA - scaling 1", type="none", xlab=c("RDA1"), ylab=c("RDA2"), xlim=c(-1,1), ylim=c(-1,1)) points(scores(spechem.physio, display="sites", choices=c(1,2), scaling=1), pch=21, col="black", bg="steelblue", cex=1.2) arrows(0,0, scores(spechem.physio, display="species", choices=c(1), scaling=1), scores(spechem.physio, display="species", choices=c(2), scaling=1), col="black",length=0) text(scores(spechem.physio, display="species", choices=c(1), scaling=1), scores(spechem.physio, display="species", choices=c(2), scaling=1), labels=rownames(scores(spechem.physio, display="species", scaling=1)), col="black", cex=0.8) arrows(0,0, scores(spechem.physio, display="bp", choices=c(1), scaling=1), scores(spechem.physio, display="bp", choices=c(2), scaling=1), col="red") text(scores(spechem.physio, display="bp", choices=c(1), scaling=1)+0.05, scores(spechem.physio, display="bp", choices=c(2), scaling=1)+0.05, labels=rownames(scores(spechem.physio, display="bp", choices=c(2), scaling=1)), col="red", cex=1) #Scaling 2 windows(title="Partial RDA scaling 2") plot(spechem.physio, scaling=2, main="Triplot partial RDA - scaling 2", type="none", xlab=c("RDA1"), ylab=c("RDA2"), xlim=c(-1,1), ylim=c(-1,1)) points(scores(spechem.physio, display="sites", choices=c(1,2), scaling=2), pch=21, col="black", bg="steelblue", cex=1.2) arrows(0,0, scores(spechem.physio, display="species", choices=c(1), scaling=2), scores(spechem.physio, display="species", choices=c(2), scaling=2), col="black",length=0) text(scores(spechem.physio, display="species", choices=c(1), scaling=2), scores(spechem.physio, display="species", choices=c(2), scaling=2), labels=rownames(scores(spechem.physio, display="species", scaling=2)), col="black", cex=0.8) arrows(0,0, scores(spechem.physio, display="bp", choices=c(1), scaling=2), scores(spechem.physio, display="bp", choices=c(2), scaling=2), col="red") text(scores(spechem.physio, display="bp", choices=c(1), scaling=2)+0.05, scores(spechem.physio, display="bp", choices=c(2), scaling=2)+0.05, labels=rownames(scores(spechem.physio, display="bp", choices=c(2), scaling=2)), col="red", cex=1)
This RDA is significant (p<0.001) as well as the two first canonical axis. Water chemistry explained 31.89% of the variance of fish species composition, physiographic covariables explained 41.53% of this variation and the unexplained variation is 26.59%. The adjusted R² of this RDA is 24.13%. The triplot looks like:
Challenge 2
Run the partial RDA of the mite environmental variables of the mite species abundances partialling out for the substrate variables (SubsDens, WaterCont and Substrate). Is the model significant? Which are the significant axes? Interpret the obtained triplot.
Challenge 2: Solution
2.3 Variation partitioning by partial RDA
Variation partitioning is a type of analysis that combines RDA and partial RDA to divide the variation of a response variable among two, three or four explanatory data sets. Variation partitioning are generally represented by Venn diagram in which the percentage of explained variance by each explanatory data set (or combination of data stets) is reported.
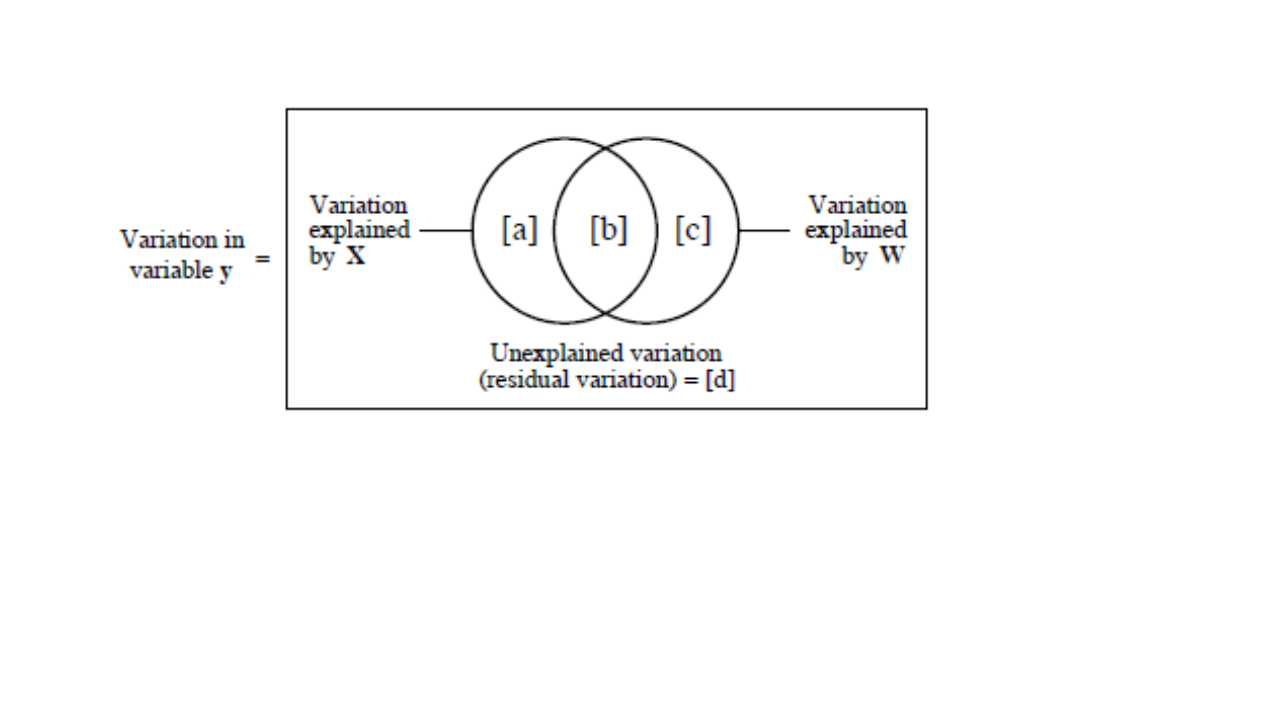
In the case of two datasets (below): - Fraction a + b +c is the explained variance by the two datasets calculated using a RDA of y by X + W. - Fraction d is the unexplained variance by the two datasets calculated using the same RDA as above. - Fraction a is the explained variance by the X data set only calculated using a partial of y by X with W as covariables. - Fraction c is the explained variance by the W data set only calculated using a partial of y by W with X as covariables. - Fraction b is calculated by subtraction, i.e. b = [a + b] + [b + c] - [a + b + c].
Venn diagram of partition of the variation of a response variable y among two sets of explanatory variables X and W (from Legendre and Legendre 2012).
Variation partitioning is thus an indicated analysis when user what to relate the abundance of species in a community to various type of environmental variables, for example abiotic vs biotic variables, large-scale versus small-scale variables, etc. In the next example, we will partition the variation of fish species composition between chemical and physiographic variables.
In R, variation partitioning is performed using the function varpart(). Venn diagrams can also be drawn using the function plot().
- | varpart()
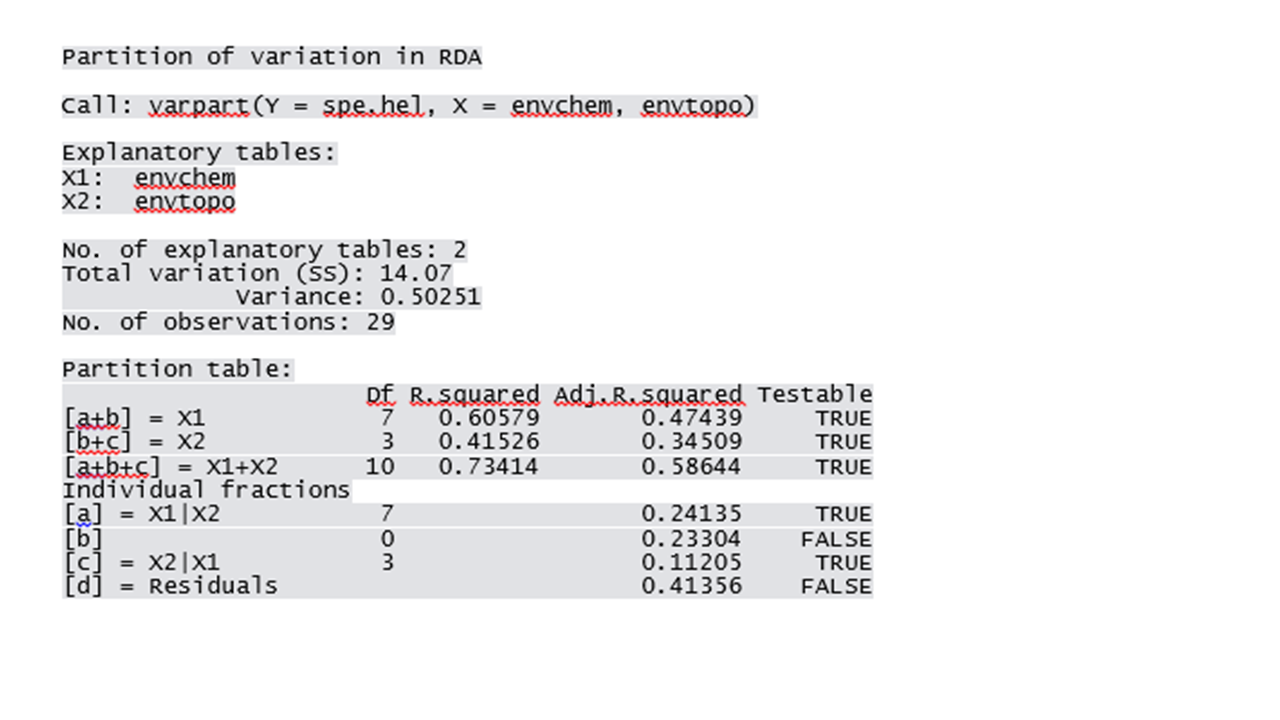
?varpart vegandocs("partitioning.pdf") #Variation partitioning with all explanatory variables spe.part.all <- varpart(spe.hel, envchem, envtopo) spe.part.all windows(title="Variation partitioning - all variables") plot(spe.part.all, digits=2)
In this case, the chemical variables explain 24.10% of the fish species composition, the physiographic variables explain 11.20% of the fish species composition and the interaction of these two types of variables explained 23.30% of the fish species composition. Note that the varpart() function also identify the fractions that can be tested for significance using the function anova.cca().
Users can also perform variation partitioning between data sets that only contain significant environmental variables:
- | varpart with significant variables
#RDA of chemistry variables spe.chem <- rda(spe.hel~., data=envchem) #Select significant chemistry variables R2a.all.chem <- RsquareAdj(spe.chem)$adj.r.squared ordiR2step(rda(spe.hel~1, data=envchem), scope= formula(spe.chem), direction= "forward", R2scope=TRUE, pstep=1000) names(envchem) (envchem.pars <- envchem[, c( 4, 6, 7 )]) #RDA with other environmental variables spe.topo <- rda(spe.hel~., data=envtopo) R2a.all.topo <- RsquareAdj(spe.topo)$adj.r.squared ordiR2step(rda(spe.hel~1, data=envtopo), scope= formula(spe.topo), direction= "forward", R2scope=TRUE, pstep=1000) names(envtopo) envtopo.pars <- envtopo[, c(1,2)] #Varpart spe.part <- varpart(spe.hel, envchem.pars, envtopo.pars) windows(title="Variation partitioning - parsimonious subsets") plot(spe.part, digits=2) #Tests of significance anova.cca(rda(spe.hel, envchem.pars), step=1000) # Test of fractions [a+b] anova.cca(rda(spe.hel, envtopo.pars), step=1000) # Test of fractions [b+c] env.pars <- cbind(envchem.pars, envtopo.pars) anova.cca(rda(spe.hel, env.pars), step=1000) # Test of fractions [a+b+c] anova.cca(rda(spe.hel, envchem.pars, envtopo.pars), step=1000) # Test of fraction [a] anova.cca(rda(spe.hel, envtopo.pars, envchem.pars), step=1000) # Test of fraction [c]
Now, the chemical variables explain 25.30% of the fish species composition, the physiographic variables explain 14.20% of the fish species composition and the interaction of these two types of variables explained 19.60% of the fish species composition. All these fractions are significant (p<0.001).
Challenge 3: Perform variation partitioning of the mite species abundances with a first dataset for the significant substrate variables (SubsDens, WaterCont and Substrate) and a second dataset for the significant other variables (Shrud and Topo). What proportion of the variation are explained by each dataset? What are the significant fractions ?
Challenge 3: Solution
3. Multivariate regression tree
Multivariate regression tree (MRT) is a constrained clustering technique. Introduced by De’ath (2002), MRTs allow the partitioning of a quantitative response matrix by a matrix of explanatory variables constraining (guiding) on where to divide the data of the response matrix. RDA and MRT are both regression techniques, the former explaining the global structure of relationships through a linear model, the latter better highlighting local structures and interactions among variables by producing a tree model.
Advantages of the MRT compared to the RDA:
- does not make assumptions about the shape of the relationships between species and environmental variables (quantitative or categorical),
- is robust in dealing with missing values
- is robust in dealing with collinearity among the explanatory variables
- is insensitive to transformations of the explanatory variables, which allows the use of raw values
- the outcome, the tree, is easy to interpret, especially to a non-scientist audience.
The MRT technique splits the data into clusters of samples similar in their species composition based on environmental value thresholds. It involves two procedures running at the same time: 1) the computation of the constrained partitioning of the data, and 2) the calculation of the relative error of the successive partitioning levels by multiple cross-validations. The function mvpart() from the package mvpart computes both the partition and the cross-validation.
A quick note on MRT terminology:
Leaf: Terminal group of sites
Node: Point where the data splits into two groups. It is characterized by a threshold value of an explanatory variable.
Branch: Each group formed by a split
1- Constrained partitioning of the data
First, the method computes all possible partitions of the sites into two groups. For each quantitative explanatory variable, the sites will be sorted in the ascending values of the variables; for categorical variables, the sites will be aggregated by levels to test all combinations of levels. The method will split the data after the first object, the second object and so on, and compute the sum of within-group sum of squared distances to the group mean (within-group SS) for the response data. The method will retain the partition into two groups minimizing the within-group SS and the threshold value/level of the explanatory variable. These steps will be repeated within the two subgroups formed previously, until all objects form their own group. In other words, when each leaf of the tree contains one object.
2- Cross-validation and pruning the tree
The mvpart function also performs a cross-validation and identifies the best predictive tree. The cross-validation procedure consists in using a subset of the objects to construct the tree, and to allocate the remaining objects to the groups. In a good predictive tree, objects are assigned to the appropriate groups. The cross-validated relative error (CVRE) is the measure of the predictive error. Without cross-validation, one would retain the number of partitions minimizing the variance not explained by the tree (i.e. the relative error: the sum of the within-group SS over all leaves divided by the overall SS of the data). This is the solution maximizing the R2 so to speak. This approach is explanatory rather than predictive.
Let’s create a multivariate regression tree on the Doubs data.
- | mvpart()
?mvpart #Prepare the data: remove “distance from source” env <- subset(env, select = -das) # Create the regression tree doubs.mrt <- mvpart(as.matrix(spe.hel) ~. ,env, legend=FALSE, margin=0.01, cp=0, xv="pick", xval=nrow(spe.hel), xvmult=100, which=4)
At this point, you will need to select the tree who's size (number of groups) is appropriate to the aim of your study from the following graph. This step requires the argument xv=“pick”. In other words, you must prune the tree by picking the best-fit tree. Indeed, a fully resolved tree is not the desirable outcome. Instead, one is usually interested in a tree including only informative partitions/groups. It is possible to have an a-priori idea of the number of potential groups to be retained as well.
The graph shows the relative error RE (in green) and the cross-validated relative error CVRE (in blue) of trees of increasing size. The red dot indicates the solution with the smallest CVRE, and the orange dot shows the smallest tree within one standard error of CVRE. It has been suggested that instead of choosing the solution minimizing CVRE, it would be more parsimonious to opt for the smallest tree for which the CVRE is within one standard error of the tree with the lowest CVRE (Breiman et al. 1984). The green bars at the top indicate the number of times each size was chosen during the cross-validation process.
This graph is interactive, which means you will have to click on the blue point corresponding your choice of tree size. Once you do so, the corresponding multivariate regression tree will appear. If you click on the orange dot, the following tree appears.
The statistics at the bottom of the figure are: the residual error (the reciprocal of the R2 of the model, in this case 43.7%), the cross-validated error, and the standard error. This tree has only two leaves separated by one node. This node splits the data into two groups at the threshold altitude value of 361.5m.
Each leaf is characterized by a small barplot showing the abundances of the species, its number of sites and its relative error.
We can compare this tree with the 10-group solution, as suggested by the CVRE criterion, or choose a solution in between, e.g. with 4 leaves to compare.
- | Compare trees
# Using the CVRE criterion doubs.mrt.cvre <- mvpart(as.matrix(spe.hel)~., env, legend=FALSE, margin=0.01, cp=0,xv="pick", xval=nrow(spe.hel), xvmult=100,which=4) # Choosing ourself the best number of partitions doubs.mrt.4 <- mvpart(as.matrix(spe.hel)~., env, legend=FALSE, margin=0.01, cp=0, xv="pick", xval=nrow(spe.hel), xvmult=100,which=4)
The 10-group solution has a high EXPLANATORY power but its predictive power (indicated by the cross-validated error) is just slightly better than that of the 2-group solution. The 4-group solution seems to be a good compromise.
More information can be obtained by looking at the summary output.
- | MRT summary
summary(doubs.mrt)
CP stands for “complexity parameter”, which is the equivalent of the variance explained by each node. The CP at nsplit 0 is the R2 of the whole tree. The summary then outlines, for each node, the best threshold values to split the data. While informative, this output is very dense. A more detailed and yet more manageable output can be generated by using the wrapper from the function MRT() of the MVPARTwrap package. Plus, this other function allows identification of discriminant species.
- | Find discriminant and indicator species
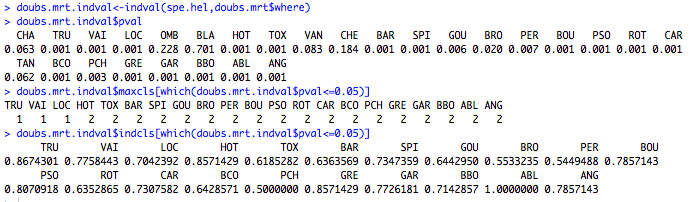
# Find discriminant species with MRT results doubs.mrt.wrap<-MRT(doubs.mrt,percent=10,species=colnames(spe.hel)) summary(doubs.mrt.wrap) # Extract indval p-values doubs.mrt.indval<-indval(spe.hel,doubs.mrt$where) doubs.mrt.indval$pval # Extract indicator species of each node, with its indval doubs.mrt.indval$maxcls[which(doubs.mrt.indval$pval<=0.05)] doubs.mrt.indval$indcls[which(doubs.mrt.indval$pval<=0.05)]
The main discriminant species of the first split are TRU, VAI and ABL. TRU and VAI contribute highly to the left leaf, and ABL is the most indicative species of the sites at lower altitude (<361.5m). This output also indicates which sites are included in each leaf.
The second part of the code allows us to test the significance of the indicator value of each species through a permutation test. For each significant indicator species, we extracted the leaf number and the indicator value. In this particular case, TRU, VAI and LOC are all significant species of the left leaf, TRU having the highest indicator value (0.867).
Challenge 4: Run the multivariate regression tree for the mite data. Select the minimum size of tree within one SE of the CVRE. What is the proportion of variance explained by this tree? How many leaves contain this tree? What are the discriminant species?
Challenge 4 - Solution
4. Linear discriminant analysis
Linear discriminant analysis (LDA) is a constrained (canonical) technique that allows you to determine how well your independent set of variables explains an a priori grouping. This grouping may have been obtained from a previous clustering analysis (see Workshop 8) or from a hypothesis (e.g. grouping is based on sites at different latitudes or different treatments). An LDA can also be used to classify new data into these pre-determined groups. You can imagine some useful applications of this technique including assessing which population a fish should be classified in based on morphology or classifying whether a new paper is a freshwater, marine or terrestrial study based on the abstract of papers in those pre-determined biomes.
LDA computes discriminant functions from standardized descriptors. These coefficients quantify the relative contributions of the (standardized) explanatory variables to the discrimination of objects. Identification functions can be computed from the original (not standardized) descriptors to classify new data into pre-determined groups. Let’s continue to work with the Doubs fish data. First we must ensure that the within-group covariance matrices of the explanatory variables are homogeneous – a condition necessary for the application of LDA.
First we want to make an a priori classification that is independent from the environmental data set. We know that there is a general relationship that indicates environmental variables change with latitude (Budyko 1969). Here we will classify our Doubs fish sites based on latitude to determine how well the environmental factors explain our latitude grouping. Our groups are determined by simply dividing the range of latitudes equally into three groups and then assigning each site to a group depending on where they fall along the divided range.
- | Load spatial data and classify Doubs fish sites based on latitude
#load spatial data to determine groups spa <- read.csv ('http://www.davidzeleny.net/anadat-r/data-download/DoubsSpa.csv', row.names = 1) spa <- spa[,-8] #View spatial data View (spa) #add site numbers numbers<-(1:30) numbers<-numbers[!numbers%in%8] spa$site<-numbers #make groups based on lattitude y<82=group1, 82<y<156=group2, y>156=group3 spa.group<-ddply(.data=spa, .variables=.(x, y, site), .fun= summarise, group = if(y <= 82) 1 else if (y <= 156) 2 else 3) #order by site spa.group<-spa.group[with(spa.group, order(site)), ]
Generally, we would first want to check that the within-group covariance matrices of the explanatory variables are homogeneous by verifying multivariate homogeneity of within-group covariance (MHV). For the purposes of this workshop we will by pass it but more information can be found in Borcard et al. (2011).
Once we run the LDA we can use the result object to determine 1. What groups the sites are classified in based on the environmental data. 2. What are the posterior probabilities of that the sites to belong to the groups. 3. The percentage of correct classification based on our latitudinal grouping.
- | Run the linear discriminant analysis (LDA)
#run LDA LDA<-lda(env,spa.group[,4]) #classification of the objects based on LDA spe.class <- predict(LDA)$class #posterior probabilities of the objects to belong to the groups spe.post <- predict(LDA)$posterior #table of prior versus predicted classifications spe.table <- table(spa.group[,4], spe.class) #proportion of correct classification diag(prop.table(spe.table, 1))
The results suggest that the environmental factors explain the first, lower latitude, group and the group 3 perfectly but only 83% of the group 2 sites were predicted correctly. What does that tell us about our classification? Perhaps there are stronger delineations in the lower and higher latitude and the group 2 is a mix of both?
Now what we have some new sites and we want to classify them based on the relationship we have established between our latitudinal grouping and environmental factors using the LDA. Using the predict() function we can load in a new matrix with sites and classify them using the LDA object.
Load in the classifyme.csv file, which contains dummy data from 5 new sites.
- | Load the classify me data and predict grouping of new data
#predicting classification of new data #read in new sites classify.me<-read.csv("classifyme.csv", header = T) #predict grouping of new data predict.group<-predict(LDA, newdata=classify.me) #give classification for each new site group.new<-predict.group$class
Our new sites, in order, have been classified in groups 1,1, 1, 3 and 3 respectively.
Challenge 5: Run an LDA for the mite env data (only first two vars) based on four latitudinal groups you create from the mite.xy data set. What group was group 2 most incorrectly grouped into? What proportion of sites was correctly classified in group 1? group 2?
Challenge 5: Solution
5. Some other useful ordination methods
- | other methods
?cca #(constrained correspondence analysis) # Constrained Correspondence Analysis (CCA) is a canonical ordination method similar to RDA that preserve # Chi-square distances among object (instead of Euclidean distances in RDA). This method is well suited for the # analysis of large ecological gradients. ?CCorA # Canonical Correlation Analysis # Canonical Correlation Analysis (CCorA) differs from RDA given that the two matrices are considered symmetric # while in RDA the Y matrix is dependent on the X matrix. The main use of this technique is to test the # significance of the correlation between two multidimensional data sets, then explore the structure of the data by # computing the correlations (which are the square roots of the CCorA eigenvalues) that can be found between # linear functions of two groups of descriptors. help(coinertia, package=ade4) # Coinertia Analysis #Coinertia Analysis (CoIA) is a symmetric canonical ordination method that is appropriate to compare pairs # of data sets that play equivalent roles in the analysis. The method finds a common space onto which the objects # and variables of these data sets can be projected and compared. Compared to CCorA, co-inertia analysis # imposes no constraint regarding the number of variables in the two sets, so that it can be used to compare # ecological communities even when they are species-rich. Co-inertia analysis is not well-suited, however, to # analyse pairs of data sets that contain the same variables, because the analysis does not establish one-to-one # correspondences between variables in the two data sets; the method does not ‘know’ that the first variable is the # same in the first and the second data sets, and likewise for the other variables. help(mfa, package=ade4) # Multiple Factorial Analysis # Multiple factor analysis (MFA) can be used to compare several data sets describing the same objects. MFA # consists in projecting objects and variables of two or more data sets on a global PCA, computed from all data # sets, in which the sets receive equal weights. # Spatial analysis can be performed using packages AEM and PCNM : http://r-forge.r-project.org/R/?group_id=195
References
Alday & Marrs (2014). A simple test for alternative states in ecological restoration: the use of principal response curves. Journal of Vegetation Science, 17, 302-311.
Borcard, Gillet & Legendre (2011). Numerical Ecology with R. Springer New York.
Breiman, L., J. H. Friedman, et al. (1984). Classification and Regression Trees. Belmont, California, USA, Wadsworth International Group.
Budyko, M.I. (1969) The effect of solar radiation variations on the climate of the Earth. Tellus, 21(5), 611-619.
Clarke & Warwick (2001). Change in Marine Communities: An Approach to Statistical Analysis and Interpretation 2nd edition. Primer-E Ltd.
De'ath, G. (2002). Multivariate regression trees : a new technique for modeling species-environment relationships. Ecology, 83(4), 1105–1117.
Gotelli & Ellison (2004). A Primer of Ecological Statistics. Sinaeuer Associates Inc., Sunderland MA.
Legendre & Legendre (2012). Numerical Ecology 3rd edition. Elsevier Science BV, Amsterdam.
Poulin, Andersen & Rochefort (2013) A new approach for tracking vegetation change after restoration: a case study with peatlands. Restoration Ecology, 21, 363-371.